File Information
| Name | libRoadRunner: High-Performance SBML Simulator |
|---|---|
| Version | v2.8.0 |
| File Size | Windows: ~380MB • macOS: ~111MB • Linux: ~147MB |
| Platforms | Windows • macOS • Linux |
| License | Open Source (Apache License 2.0) |
| Official Repository | GitHub – sys-bio/roadrunner |
| Official Site | libroadrunner |
Table of contents
Description
libRoadRunner is an open-source, high-performance simulation engine for systems biology models written in SBML (Systems Biology Markup Language). Built in C/C++ and powered by LLVM JIT compilation, it delivers lightning-fast performance for large-scale biological simulations. Researchers, scientists, and developers use libRoadRunner for metabolic modeling, steady-state analysis, and time-dependent simulations.
This versatile simulator can be integrated into Python, C++, or Julia environments through official bindings. It’s ideal for parameter optimization, model analysis, and computational biology pipelines running on both personal systems and HPC clusters.
Whether you’re simulating biological pathways, running control analyses, or building complex systems models, libRoadRunner provides a robust, efficient, and modular backend. It’s available as open source under the Apache License 2.0, ensuring complete flexibility for academic and commercial use.
Features of LibRoadRunner
| Feature | Description |
|---|---|
| High Performance | Uses LLVM JIT compilation to generate optimized machine code for extremely fast simulations. |
| Multi-language Support | Offers bindings for C/C++, Python, and Julia, allowing integration across various workflows. |
| Comprehensive SBML Support | Compatible with SBML Levels 2–3, supporting most model features except algebraic rules and DDEs. |
| Flexible Simulation Modes | Supports steady-state, time-dependent, frequency domain, and control analysis simulations. |
| Plugin System | Extend functionality using plugins like the Levenberg-Marquardt optimizer. |
| Built-in Analysis Tools | Access to eigenvalues, eigenvectors, Jacobian matrices, and structural matrices for deep model insights. |
| Cross-Platform | Available for Windows, macOS (Intel + Apple Silicon), and major Linux distributions. |
| Self-contained Binaries | Includes all dependencies — no external libraries needed except ncurses on Linux. |
Screenshots
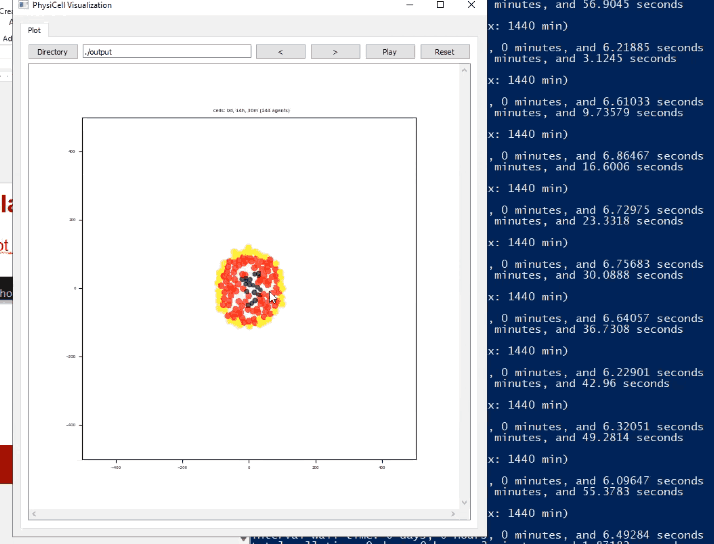
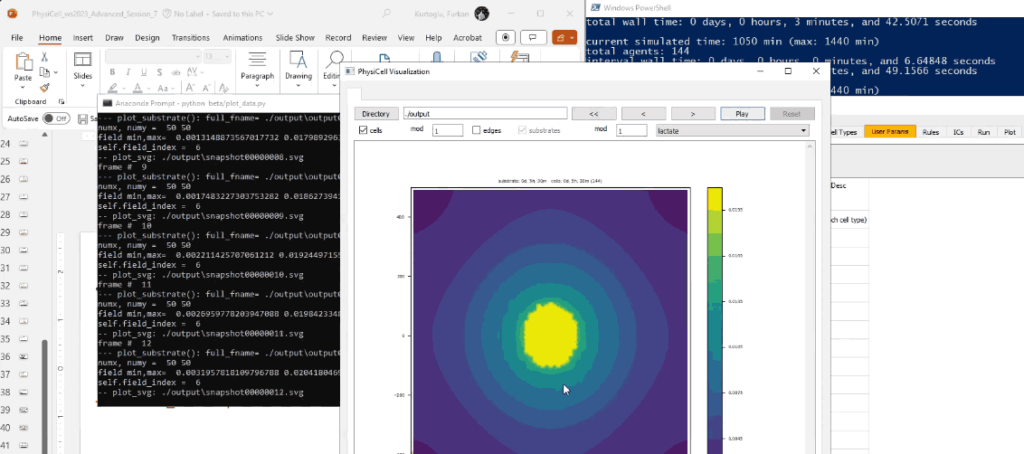
System Requirements
| Component | Minimum Requirement |
|---|---|
| Operating System | Windows 10/11 • macOS 12+ (Intel or ARM) • Linux (Ubuntu 20.04+, CentOS 8, etc.) |
| Processor | 64-bit CPU with SSE2 support |
| RAM | 4 GB minimum (8 GB recommended) |
| Storage | 500 MB available space |
| Dependencies (Linux) | ncurses library |
| Python Integration | Python 3.10 or higher (for wheel installs) |
How to Install libroadrunner??
Windows / macOS / Linux
- Download the corresponding binary (
.zip) for your platform. - Extract the archive to your preferred directory.
- Run the executable or integrate libRoadRunner as a library in your preferred language.
- (For Python users) Install directly using:
pip install libroadrunner
Download librunnerroad High-Performance SBML Simulation Library
Conclusion
libRoadRunner stands as one of the most advanced open-source simulation engines for systems biology and computational modeling. Its exceptional performance, powered by LLVM JIT compilation, makes it ideal for researchers, developers, and scientists working with SBML models. With native support for Windows, macOS, and Linux, seamless Python and C++ integration, and an extensive plugin ecosystem, it bridges the gap between accuracy, speed, and flexibility in biological simulations.





